-Search query
-Search result
Showing 1 - 50 of 426 items for (author: fan & sl)

EMDB-43508: 
Structure of a bacterial gasdermin small oval pore assembly
Method: single particle / : Johnson AG, Mayer ML, Kranzusch PJ

EMDB-43509: 
Structure of a bacterial gasdermin medium oval pore assembly
Method: single particle / : Johnson AG, Mayer ML, Kranzusch PJ

EMDB-43510: 
Structure of a bacterial gasdermin large oval pore assembly
Method: single particle / : Johnson AG, Mayer ML, Kranzusch PJ

EMDB-43511: 
Structure of a bacterial gasdermin double pore assembly
Method: single particle / : Johnson AG, Mayer ML, Kranzusch PJ
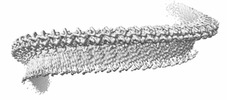

EMDB-43513: 
Structure of a bacterial gasdermin slinky-like oligomer from a heterogeneous sample
Method: single particle / : Johnson AG, Mayer ML, Kranzusch PJ
EMDB-37356: 
Cryo-EM structure of the GPR101-Gs complex
Method: single particle / : Sun JP, Gao N, Yu X, Wang GP, Yang F, Wang JY, Yang Z, Guan Y

EMDB-37357: 
Cryo-EM structure of the AA-14-bound GPR101-Gs complex
Method: single particle / : Sun JP, Yu X, Gao N, Yang F, Wang JY, Yang Z, Guan Y, Wang GP

EMDB-37358: 
Cryo-EM structure of the AA14-bound GPR101 complex
Method: single particle / : Sun JP, Yu X, Gao N, Yang F, Wang JY, Yang Z, Guan Y, Wang GP

EMDB-29227: 
cryoEM structure of a broadly neutralizing antibody STI-9167
Method: single particle / : Bajic G

EMDB-42250: 
Varicella-zoster virus glycoprotein B; H527P prefusion class I.
Method: subtomogram averaging / : Zhou M, Oliver SL, Muyuan C, Vollmer B

EMDB-42251: 
Varicella-zoster virus glycoprotein B; H527P prefusion mutant class II.
Method: subtomogram averaging / : Zhou M, Oliver SL, Muyuan C, Vollmer B

EMDB-37716: 
Human calcium-sensing receptor bound with cinacalcet in detergent
Method: single particle / : Ling SL, Meng XY, Tian CL

EMDB-37724: 
Human calcium-sensing receptor(CaSR) bound to cinacalcet in complex with Gq protein
Method: single particle / : Ling SL, Meng XY, Tian CL

EMDB-28537: 
cryoEM structure of a broadly neutralizing anti-SARS-CoV-2 antibody STI-9167
Method: single particle / : Bajic G

PDB-8eqf: 
cryoEM structure of a broadly neutralizing anti-SARS-CoV-2 antibody STI-9167
Method: single particle / : Bajic G

EMDB-27813: 
Structure of monomeric LRRK1
Method: single particle / : Reimer JM, Mathea S, Chatterjee D, Knapp S, Leschziner AE

EMDB-27814: 
Local refinement around RCKW of LRRK1
Method: single particle / : Reimer JM, Mathea S, Knapp S, Leschziner AE

EMDB-27815: 
Local refinement of LRRK1 around the ROC-COR-kinase domains
Method: single particle / : Reimer JM, Mathea S, Knapp S, Leschziner AE

EMDB-27816: 
Local refinement around kinase and WD40 domains of LRRK1
Method: single particle / : Reimer JM, Mathea S, Knapp S, Leschziner AE

EMDB-27817: 
Structure of dimeric LRRK1
Method: single particle / : Reimer JM, Lin YX, Leschziner AE

EMDB-27818: 
Symmetry expansion of dimeric LRRK1
Method: single particle / : Reimer JM, Lin YX, Leschziner AE

EMDB-16157: 
Map of GroEL:GroES-(ADP) complex plunge frozen 200 ms after reaction initiation with ATP
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16091: 
Cryo-EM structure of beta-galactosidase at 3.3 A resolution plunged 5 ms after mixing with apoferritin
Method: single particle / : Torino S, Dhurandhar M, Efremov R

EMDB-16092: 
Cryo-EM structure of beta-galactosidase at 2.9 A resolution plunged 205 ms after mixing with apoferritin
Method: single particle / : Torino S, Dhurandhar M, Efremov R

EMDB-16093: 
Cryo-EM structure of mouse heavy-chain apoferritin at 2.1 A plunged 5ms after mixing with b-galactosidase
Method: single particle / : Torino S, Dhurandhar M, Efremov R

EMDB-16094: 
Cryo-EM structure of mouse heavy-chain apoferritin at 2.7 A plunged 35ms after mixing with b-galactosidase
Method: single particle / : Torino S, Dhurandhar M, Efremov R

EMDB-16095: 
Cryo-EM structure of mouse heavy-chain apoferritin at 2.2 A plunged 205ms after mixing with b-galactosidase
Method: single particle / : Torino S, Dhurandhar M, Efremov R

EMDB-16097: 
Cryo-EM structure of beta-galactosidase at 3.2 A resolution plunged 35 ms after mixing with apoferritin
Method: single particle / : Torino S, Dhurandhar M, Efremov R

EMDB-16099: 
GroEL:GroES-ATP complex under continuous turnover condirions
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16100: 
Structure of GroEL-ATP complex plunge frozen 200 ms after reaction initiation
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16102: 
Structure of GroEL-nucleotide complex in ADP-like conformation plunged 13 ms after mixing with ATP
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16106: 
Structure of the GroEL-ATP complex plunge-frozen 50 ms after mixing with ATP
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16107: 
Structure of the GroEL(ATP7/ADP7) complex plunged 13 ms after mixing with ATP
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16108: 
Structure of GroEL-nucleotide complex in ADP-like conformation plunged 50 ms after mixing with ATP
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16109: 
Structure of the GroEL(ATP7/ADP7) complex plunged 50 ms after mixing with ATP
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16115: 
Structure of the GroEL-ATP complex plunge-frozen 13 ms after mixing with ATP
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16116: 
Structure of GroEL:GroES-ATP complex plunge frozen 200 ms after reaction initiation
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16117: 
Structure of GroEL:GroES-ATP complex under continuous turnover conditions
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16118: 
Structure of GroEL-ATP complex under continuous turnover conditions
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16119: 
Structure of GroEL:GroES complex exhibiting ADP-conformation in trans ring obtained under the continuous turnover conditions
Method: single particle / : Dhurandhar M, Torino S, Efremov R

EMDB-16125: 
Structure of GroEL:GroES-ATP complex plunge frozen 200 ms after reaction initiation
Method: single particle / : Dhurandhar M, Efremov R, Torino S

EMDB-16154: 
Map of the GroEL-ES-ATP complex plunge-frozen 50 ms after mixing with ATP
Method: single particle / : Dhurandhar M, Torino S, Efremov R

PDB-8bk7: 
Cryo-EM structure of beta-galactosidase at 3.3 A resolution plunged 5 ms after mixing with apoferritin
Method: single particle / : Torino S, Dhurandhar M, Efremov R

PDB-8bk8: 
Cryo-EM structure of beta-galactosidase at 2.9 A resolution plunged 205 ms after mixing with apoferritin
Method: single particle / : Torino S, Dhurandhar M, Efremov R

PDB-8bk9: 
Cryo-EM structure of mouse heavy-chain apoferritin at 2.1 A plunged 5ms after mixing with b-galactosidase
Method: single particle / : Torino S, Dhurandhar M, Efremov R

PDB-8bka: 
Cryo-EM structure of mouse heavy-chain apoferritin at 2.7 A plunged 35ms after mixing with b-galactosidase
Method: single particle / : Torino S, Dhurandhar M, Efremov R

PDB-8bkb: 
Cryo-EM structure of mouse heavy-chain apoferritin at 2.2 A plunged 205ms after mixing with b-galactosidase
Method: single particle / : Torino S, Dhurandhar M, Efremov R

PDB-8bkg: 
Cryo-EM structure of beta-galactosidase at 3.2 A resolution plunged 35 ms after mixing with apoferritin
Method: single particle / : Torino S, Dhurandhar M, Efremov R

PDB-8bkz: 
GroEL:GroES-ATP complex under continuous turnover conditions
Method: single particle / : Dhurandhar M, Torino S, Efremov R

PDB-8bl2: 
Structure of GroEL-ATP complex plunge frozen 200 ms after reaction initiation
Method: single particle / : Dhurandhar M, Torino S, Efremov R
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
